Welcome to OpenSILEX
An Open Source information system.
 Try on Sandbox
User Documentation (en) / Guide Utilisateur (fr)
Try on Sandbox
User Documentation (en) / Guide Utilisateur (fr)
 Try on Sandbox
User Documentation (en) / Guide Utilisateur (fr)
Try on Sandbox
User Documentation (en) / Guide Utilisateur (fr)
OpenSILEX is a set of interoperable software modules (structuring, visualization, analysis, reproducibility, etc.) which allows the creation of scientific information systems dedicated to the exploitation of large-scale data in agronomy.
OpenSILEX instances are used on a daily basis by around ten research units around the world, technical institutes and partnerships (e.g., Terre-Inovia, IFV, FUI Pilotype, Moët-et-Chandon, Lallemand), for example :
Thanks to concrete features for sharing and reusing data, OpenSILEX greatly promotes the development of open science and the creation of value around data.
The PHIS instance, for example, has established itself as an international benchmark for the structuring and exploitation of large-scale data produced by high-throughput phenotyping with the Phenome or EMPHASIS platforms.
The work around OpenSilex is based on strong collaborations (INRIA, University of Montpellier, I3S CNRS, WUR (NL), CNR (It), Univ Tokyo).
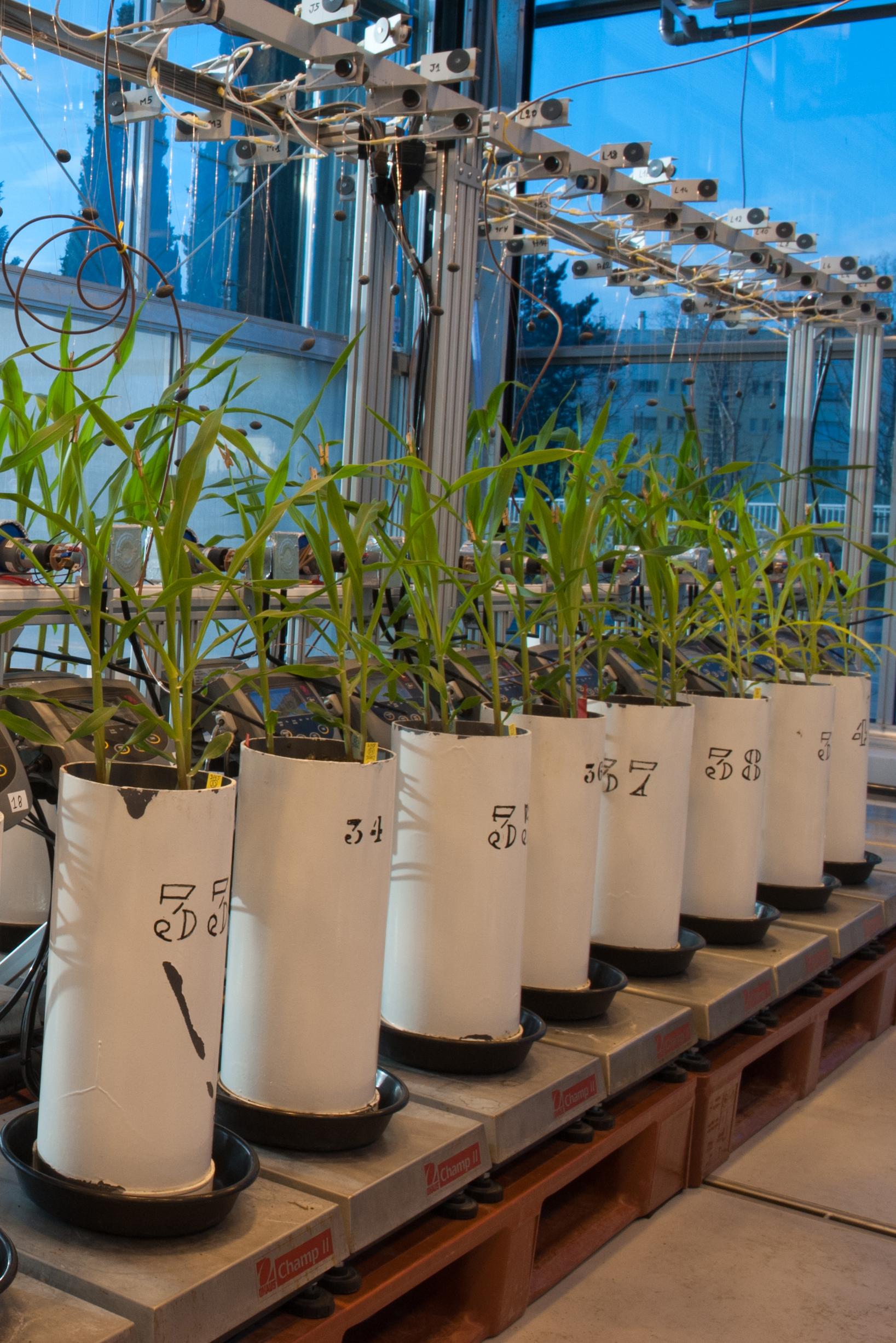

OpenSILEX is an ontology-driven Information System designed for life science data.
OpenSILEX main features
The Service API allows integration of data from external databases and resources, export to computing and modelling platforms and integration of phenomic data into other systems.
A Web user interface allows exploration of data through interactive and user-friendly menus and tools.
OpenSILEX is available as an open source software under a GNU Affero General Public License version 2. Source code, user and developer documentation are available at github.com/OpenSILEX.
The use of URIs allows standardized, unique and unambiguous identification of objects involved in experiments. Generate your own objects uri: URI Generator.
Datasets are enriched with knowledge and metadata enabling the reuse of data and meta-analyses.
Tracking features based on object identifcation and events allows following the different objects involved in experiment in time and space.
OpenSILEX allows to store, organise and manage heterogenous multi-deminsional data originating from multiple sources.
An inference engine based on the semantics and rules represented in the ontologies enables advanced queries, thereby linking the knowledge stored in the Triple Store to the information distributed among the different storage systems.
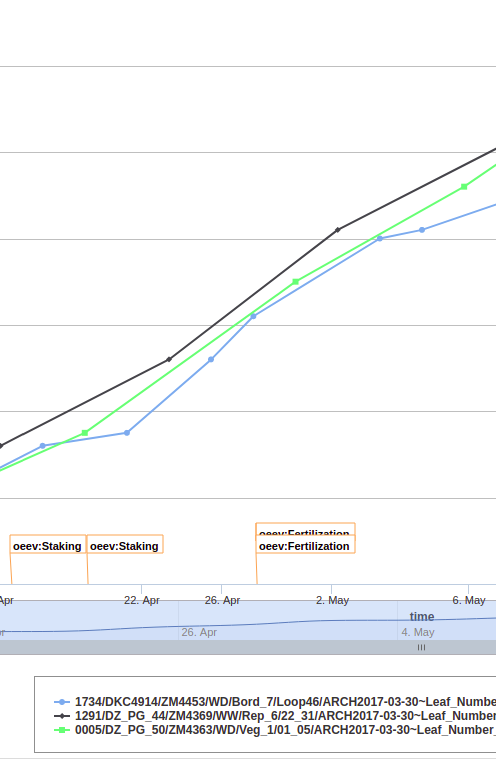
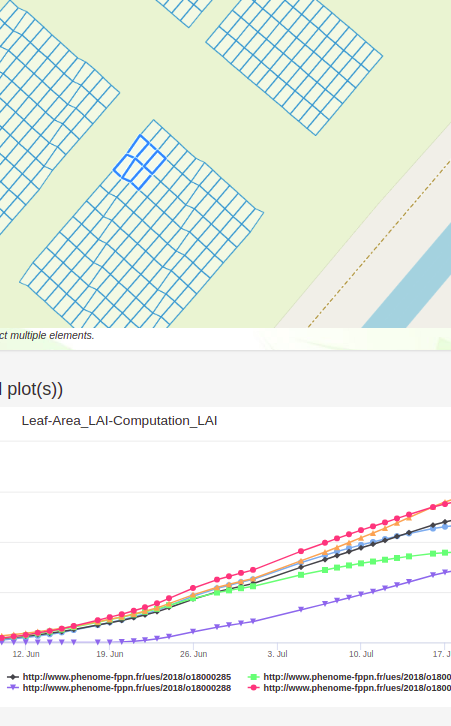
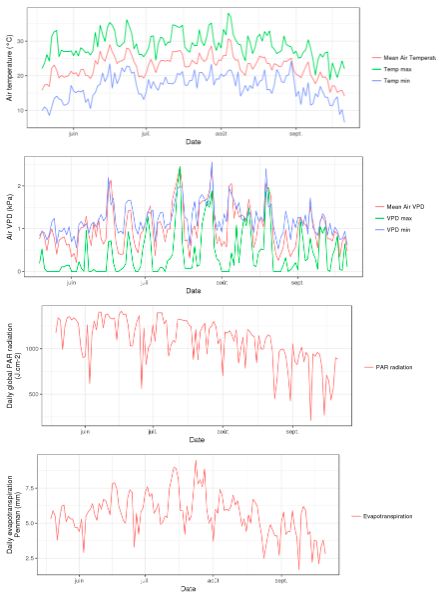
Advanced visualization features allow displaying images, dynamic graphs of static or time courses of variables.
Different tools allow advanced data querying and computing features including data analysis tools and integration of workflows.
OpenSILEX is a hybrid system based on different storage databases (RDF4J, MongoDB, iRODS).


















2, place Pierre Viala
Campus La Gaillarde, bat. 29
34060 Montpellier Cedex 2
FRANCE